Channels
Special Offers & Promotions
Integrated DNA Technologies Launches xGen Universal Blocking Oligos
Enhancing the performance of target capture experiments
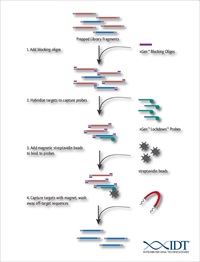 Integrated DNA Technologies (IDT), the world leader in oligonucleotide synthesis, has launched the latest addition to its portfolio of next generation sequencing (NGS) products, xGen® Universal Blocking Oligos. By incorporating these oligos into target capture experiments, users can decrease the complexity and increase throughput of their target capture experiment without degradation of performance.
Integrated DNA Technologies (IDT), the world leader in oligonucleotide synthesis, has launched the latest addition to its portfolio of next generation sequencing (NGS) products, xGen® Universal Blocking Oligos. By incorporating these oligos into target capture experiments, users can decrease the complexity and increase throughput of their target capture experiment without degradation of performance.
Blocking oligos are commonly used to enhance the performance of capture experiments, through the prevention of cross-hybridization between adapter sequences. Adapters ligated to the DNA library before enrichment can hybridize to each other and cause an increase in off-target reads. Blocking oligos bind to these adapters to prevent non-specific binding. However, when multiple samples are processed simultaneously, each adapter requires a unique blocking oligo, increasing the complexity of the experiment. xGen Universal Blocking Oligos are able to bind to all indexed adapters using a single oligonucleotide sequence, removing the need for multiple oligos. As a result, researchers can more readily multiplex their experiments for cost-effective enrichment of 1000s of samples.
Universal Blocking Oligos join xGen Lockdown® Probes, which are individually synthesized 60-120 nucleotide 5’-biotinylated capture oligos, as part of a growing offering of NGS products. Each probe is designed to enable high depth sequencing and analysis of specific regions of the genome. Whether used with custom capture panels, or for supplementing existing panels, xGen Lockdown Probes provide approximately 5000-fold enrichment, with high coverage over targeted regions and minimal GC bias.
xGen Target Capture products are ideal for users who want to focus on relevant genomic regions or transcripts of interest, or to detect single nucleotide polymorphisms, insertions/deletions, and other genetic variants using NGS.
more about xGen Lockdown Probes
Media Partners